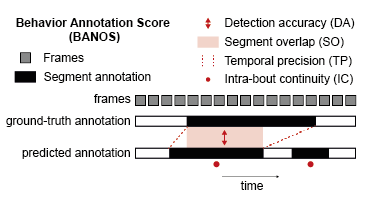
BANOS is a set of metrics for evaluating algorithmic behavior annotations against a ground truth (typically human annotations). It is designed for ethology and computer vision workflows where researchers annotate behaviors in video data.
Traditional frame-based metrics (precision/recall/F1) capture detection accuracy but miss important temporal qualities: does the predicted segment overlap well with the ground-truth bout? Does the prediction start and end at the right time? Is the prediction internally consistent, or does it flicker on and off inside the bout?
BANOS addresses these questions with four complementary metrics.
-
Detection Accuracy (DA) — Precision, Recall, and F1 score over matched behavioral segments (TP/FP/FN at the bout level).
-
Segment Overlap (SO) — Temporal Intersection over Union (tIoU) between each matched predicted segment and its ground-truth counterpart.
-
Temporal Precision (TP) — Penalizes start/end time errors:
1 / (1 + |start deviation| + |end deviation|). -
Intra-bout Continuity (IC) — Measures label stability inside each ground-truth bout:
1 - (label switches / bout length).
pip install banosRequires Python 3.9+. Core dependency: pandas. Optional: scipy (for
matching='optimal').
import banos
import pandas as pd
# Load annotations as DataFrames (rows=frames, columns=behaviors, values=0/1)
pred = pd.read_csv("machine_annotations.csv")
gt = pd.read_csv("human_annotations.csv")
# Score a single recording
metrics = banos.score(pred, gt)
# {'behavior1': {'precision': 0.9, 'recall': 0.85, 'f1_score': 0.87,
# 'so': 0.76, 'tp': 0.82, 'ic': 0.91}, ...}
# Score multiple recordings
group_metrics, overall_metrics = banos.score({
'recording_1': (pred_1, gt_1),
'recording_2': (pred_2, gt_2),
})banos.score() returns:
- Single recording:
dict[behavior → dict[metric → float]] - Multiple recordings:
(per-recording metrics, overall averaged metrics)
Absent-behavior scoring (v0.2.0): when a behavior has no ground-truth bouts, BANOS checks whether the machine also predicted nothing — if so, all metrics = 1.0 (correct absence); if the machine predicted bouts that don't exist in GT, all metrics = 0.0 (false detection). This is excluded from NaN-averaging correctly.
| Resource | Description |
|---|---|
demo/quickstart.py |
Minimal Python example — score annotations in two lines |
demo/quickstart.m |
Minimal MATLAB example |
demo/tutorial_python.ipynb |
Full Python tutorial: classic BANOS usage + frame F1 vs BANOS comparison |
demo/tutorial_matlab.m |
Full MATLAB tutorial mirroring the Python notebook |
demo/tutorial_calms21_task1.ipynb |
Human vs machine: CalMS21 Task 1 baseline, F1 + BANOS (Python only) |
By default, predicted segments are matched to ground-truth bouts greedily (by overlap). For a globally optimal assignment, use the Hungarian algorithm:
metrics = banos.score(pred, gt, matching='optimal')Requires scipy (pip install scipy).
from banos import preprocess_data, calculate_banos_for_each_file, aggregate_metrics
data_dict = {
'file1': (pred_df_1, gt_df_1),
'file2': (pred_df_2, gt_df_2),
}
preprocessed, dropped = preprocess_data(data_dict)
per_file_metrics = calculate_banos_for_each_file(preprocessed)
group_metrics, overall = aggregate_metrics(per_file_metrics)A MATLAB implementation is provided in matlab/BANOS/. Usage mirrors the Python API.
% Add toolbox to path
run('matlab/BANOS/setup.m');
% Score a single recording (pred/gt = cell arrays: row 1 = headers, rows 2+ = binary data)
metrics = BANOS_score(pred, gt);
% Score multiple recordings
dataStruct.Recording_1 = {pred1, gt1};
dataStruct.Recording_2 = {pred2, gt2};
[groupMetrics, overallMetrics] = BANOS_score(dataStruct);See demo/tutorial_matlab.m for a full walkthrough.
Also available on MATLAB File Exchange.
BANOS is functionally complete by design. We welcome bug reports and documentation improvements. See CONTRIBUTING.md for details on how to report issues, submit fixes, and the project's maintenance lifecycle.
To report a security vulnerability, see SECURITY.md — please do not use public GitHub Issues for security reports.
If you use BANOS, please cite:
Chindemi G., Bellone C., Girard B. From eye to AI: studying rodent social behavior in the era of machine learning. arXiv:2508.04255 (2025). https://doi.org/10.48550/arXiv.2508.04255
@article{chindemi2025eye,
title = {From eye to {AI}: studying rodent social behavior in the era of machine learning},
author = {Chindemi, Giuseppe and Bellone, Camilla and Girard, Benoit},
journal = {arXiv preprint arXiv:2508.04255},
year = {2025},
doi = {10.48550/arXiv.2508.04255}
}See CHANGELOG.md for release history.
MIT — see LICENSE.