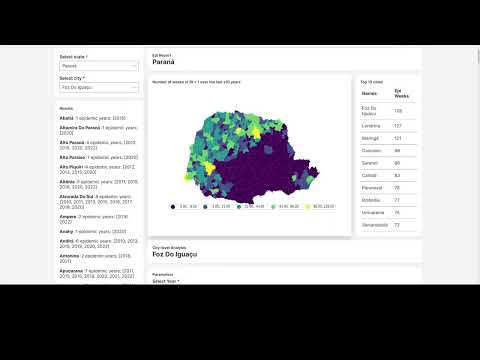
This H2O Wave application analyzes the expansion of dengue and chikungunya in Brazil using up-to-date incidence data from Infodengue.
You can set up your development environment using either Python venv (simplest) or Conda/Mamba (best for handling system dependencies like GEOS or libpq).
This approach is best if you already have the necessary system libraries (or if your OS handles them automatically via wheels).
# Create and activate virtual environment
python -m venv .venv
source .venv/bin/activate
# Install Poetry and dependencies
pip install poetry
poetry install --with devUse this if you encounter issues with libraries like geopandas or psycopg2.
- Miniforge or Anaconda installed
# Install base environment (Python + system libs + poetry)
mamba env create -f conda/env-base.yaml
conda activate episcanner
# Install project dependencies via Poetry
poetry install --with devpython ./epi_scanner/management/cli.py [disease]Then start the app:
wave run --no-reload --no-autostart epi_scanner.appMakim is used to standardize helper commands for local, CI, and production container workflows.
# Start/build the container stack for the Wave app
makim episcanner.deploy --profile dev --service wavemakim episcanner.container-wait --service wavemakim episcanner.down --profile dev --service waveTip: You can replace
devwithciorproddepending on your target environment.
The data update is now generated by the epi_scanner-downloader tool.
For more information, please see the epi_scanner-downloader repository.